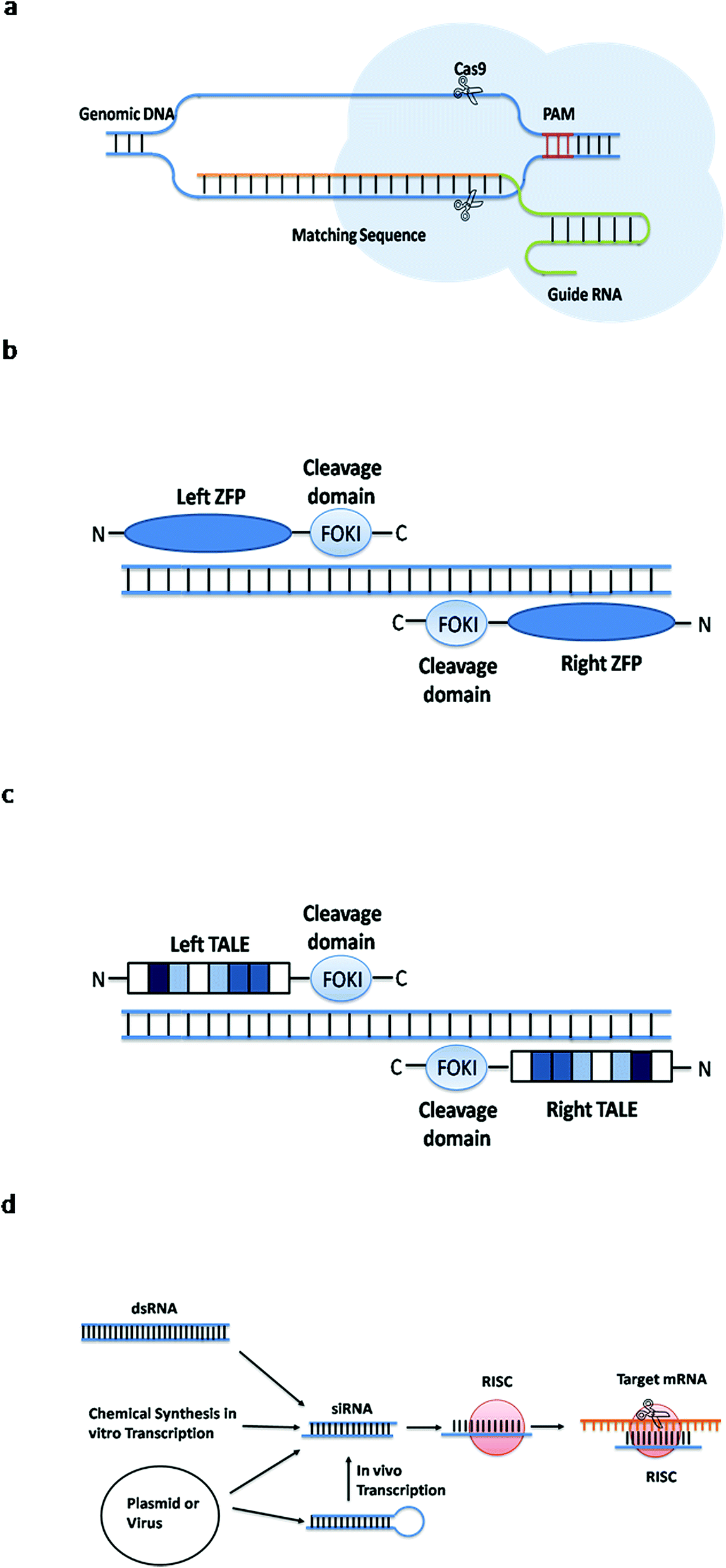
These have been extensively investigated in eukaryote model organisms such as mice and monkeys, or human cancer cell lines. Since their emergence, CRISPR-based methods have constantly been connected to concerns about off-targeting and other adverse effects. Adapting a plasmid to a novel target can be as simple as exchanging 17–20 bp within the guide RNA (gRNA). The main advantage of CRISPR/Cas over similar tools like TALENS or zinc finger nucleases is the simplicity of reprogramming it to a specific target. These include homologous recombination (HR), non-homologous end joining (NHEJ), and microhomology-mediated end joining (MMEJ). It facilitates the introduction of targeted double-strand DNA breaks (DSBs), which can trigger multiple DNA repair mechanisms in the cell. This fantastic tool was derived from a natural defense mechanism in bacteria and archaea. Many of the toolboxes recently developed around CRISPR/Cas (clustered regularly interspaced short palindromic repeats/CRISPR-associated protein) proved to be more efficient than conventional genetic tools. The practical implementation of the optimal production strain, however, is regularly hampered by the demanding process of genetic engineering. Genetic engineering of microbes has led towards promising developments in the production of pharmaceutical products, alternative fuels, and meat alternatives, among other important proteins and bulk chemicals. The identified de novo off-target mutations were rare and randomly distributed, with no apparent connection to unspecific CRISPR/Cas9 off-target binding sites. Targeting of two loci within a putatively non-essential region led to a truncation of chromosome 3 at the target locus in multiple cases, causing the deletion of 20 genes and several ribosomal DNA repeats. We also observed an increase of on-target structural variants in double target experiments as compared to single target experiments. The analysis of on-target mutations showed an unexpectedly high number of large deletions and chromosomal rearrangements at the CRISPR target loci. Furthermore, we examined the extent of possible large deletions by targeting a large genomic region, which is likely to be non-essential. We compared the outcomes of single target CRISPR transformations to double target experiments. To characterize on- and off-target mutations caused by CRISPR/Cas9 followed by non-homologous end joining repair, we chose a diverse set of CRISPR/Cas targets and conducted whole genome sequencing on 146 CRISPR/Cas9 engineered single colonies. Therefore, genome editing using CRISPR/Cas represents a simple and efficient alternative. To our knowledge this is the first example of gene therapy for an autosomal dominant gain-of-function disease using a disease allele inactivation strategy in place of the less efficient disease allele repair approach.The industrially important non-conventional yeast Komagataella phaffii suffers from low rates of homologous recombination, making site specific genetic engineering tedious. The WHIM allele-inactivated HSCs retained long-term pluripotency and selective hematopoietic reconstitution advantages. Then, through in vivo natural selection, WHIM allele-inactivated cells were enriched over wildtype allele-inactivated cells. First, one copy of Cxcr4 in HSPCs was inactivated in vitro by CRISPR/Cas9 editing with a single guide RNA (sgRNA) that does not discriminate between WHIM and wildtype Cxcr4 alleles. We have now developed a two-step preclinical protocol of autologous hematopoietic stem and progenitor cell (HSPC) transplantation towards this goal. We previously reported that Cxcr4-haploinsufficient ( Cxcr4 +/o) hematopoietic stem cells (HSCs) have a strong selective advantage for durable hematopoietic reconstitution over wildtype ( Cxcr4 +/+) and WHIM ( Cxcr4 +/w) HSCs, and that a WHIM patient was spontaneously cured by chromothriptic deletion of the disease allele in an HSC, suggesting that WHIM allele inactivation through gene editing may be a safe genetic cure strategy for the disease. WHIM syndrome is an autosomal dominant immunodeficiency disorder caused by gain-of-function mutations in chemokine receptor CXCR4 that promote severe panleukopenia due to bone marrow (BM) retention of mature leukocytes.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
